Assembline
Assembline is an assembly line of macromolecular assemblies!
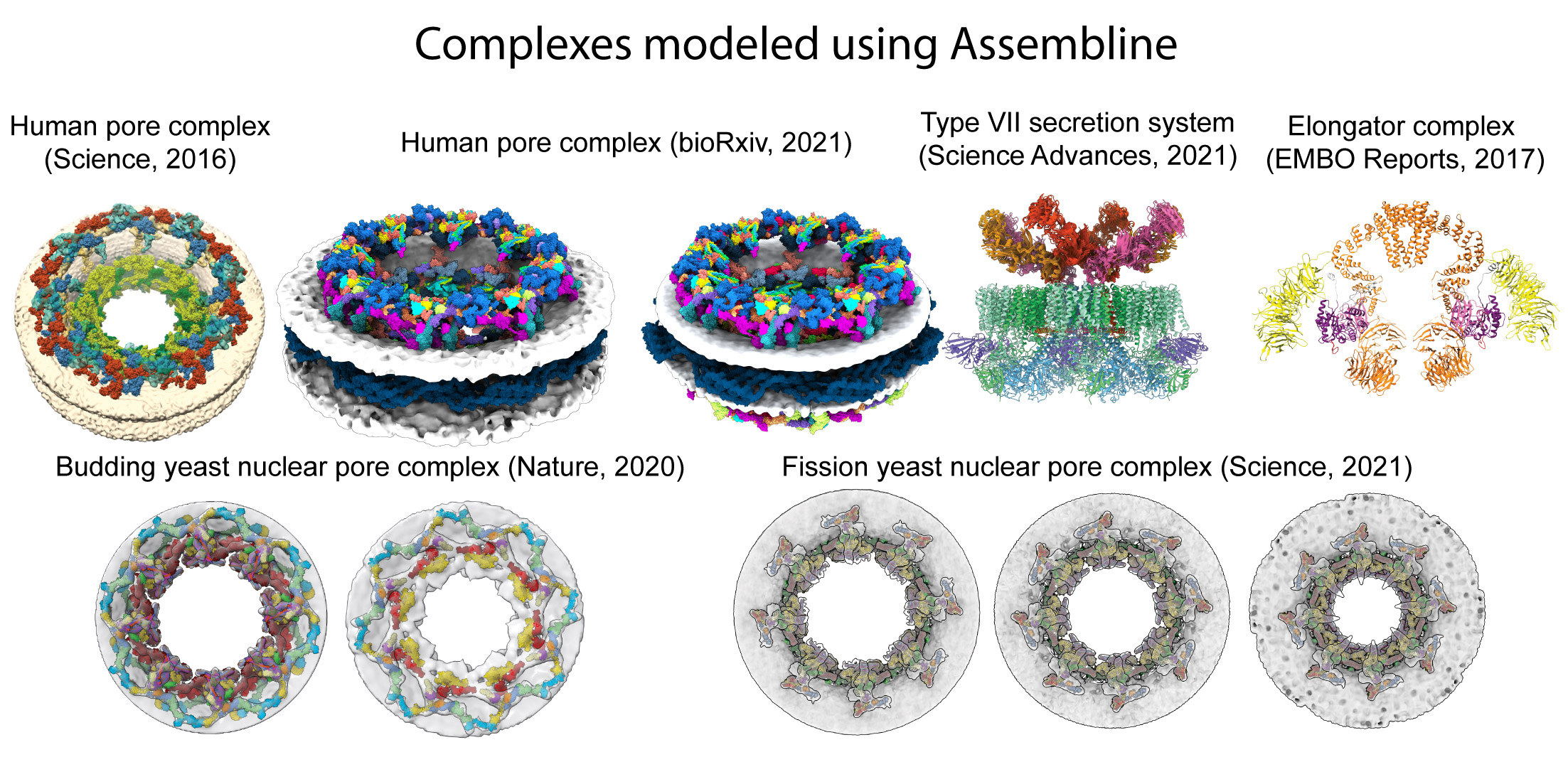
Assembline is a multi-step protocol for integrative structural modeling of macromolecular complexes based on electron microscopy, cross-linking mass spectrometry and other data. The protocols is based on our Xlink Analyzer and external software: Integrative Modeling Plafform (IMP), Python Modeling Interface (PMI), UCSF Chimera, and imp-sampcon. Comparing to other methods, Assembline enables efficient sampling of conformational space through a multi-step procedure, provides new modeling restraints, and includes a unique configuration system for setting up the modelling project.
Code
Tutorials
How to cite
If you use this software, please cite it along the dependencies it uses internally, at least:
- Rantos V., Karius K., and Kosinski J. Integrative structural modeling of macromolecular complexes using Assembline, Nature Protocols, 2021
- Integrative Modeling Plafform (IMP)
- Python Modeling Interface (PMI)
- UCSF Chimera
- Kosinski, J., et al. Xlink Analyzer: Software for analysis and visualization of cross-linking data in the context of three-dimensional structures. J. Struct. Biol. (2015)
- imp-sampcon tool used for analysis
Contact:
Please do not hesitate to send requests, comments and bug reports to:
Jan Kosinski, jan.kosinski@embl.de